S. pombe Haploid Deletion Mutant Set ver 6.0 Life Time (3,497 strains)

Yeast genome-wide knockout libraries
본 제품은 유전자 적중 방법으로 제조된 세계 유일의 분열 효모 유전자 결손 라이브러리로서 (Nature Biotech., 2010), S.pombe haploid 3,497 deletion mutants로 구성되어 있으며, 유전자 결손 시 나타나는 phenotype을 바탕으로 유전자 기능 분석 뿐 아니라, 약효 반응 분석을 통한 작용 기전 및 신약 타겟 규명 연구에도 적용 가능 합니다.
특장점
-
S.pombe 각각의 유전자의 deletion mutants로 구성
-
포유 동물과 생리학적인 프로세스 유사
-
인체 암 유전자와 30% 이상의 상동성
-
세포 주기가 짧아 (~3h) 분자생물학적인 작용 기전 및 pathway 연구에 용이
-
열성 형질에 따른 phenotype 분석이 가능
-
functional complementation을 바탕으로 알려지지 않은 유전자 기능 분석이 가능
-
살아있는 세포 기반의 대용량 약물 작용점 스크리닝에 이용 가능
개요
분열효모 게놈 적중 라이브러리는 바이오니아, 한국생명공학연구원 (KRIBB) 및 영국 왕립 암연구소 (Cancer Research UK)의 공동 연구로 개발되었으며(Nature Biotech., 2010), 바이오니아가 독점 사업권을 가지고 있습니다. 분열효모 게놈 적중 라이브러리는 유전체 수준의 유전자 기능 분석, 약물 작용점 규명 및 검증 연구, 총괄적인 세포 기능 연구를 대상으로 한 강력한 연구 수단입니다. 또한, 유전자 발현 및 합성치사(synthetic lethal) 분석 등과 같은 유전자 기능 분석 및 약물 스크리닝에 사용할 수 있습니다.
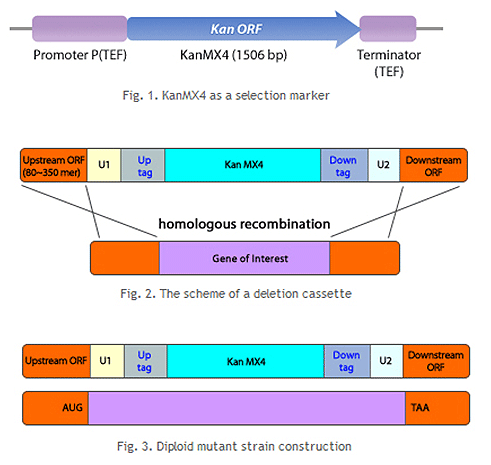
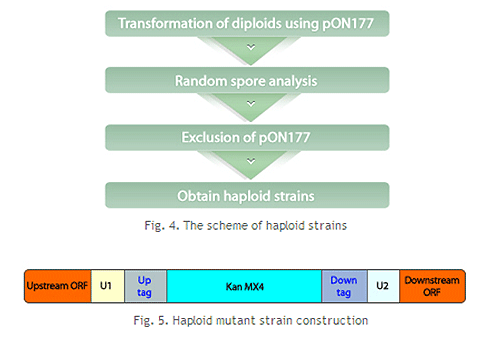
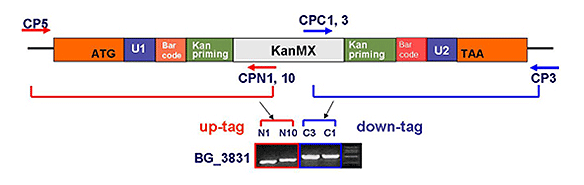
분열효모 게놈 적중 라이브러리는 상동 재조합(homologous recombination)을 통해 각 표적 유전자를 selective marker (KanMX4)를 포함한 카세트로 치환하는 방법으로 제조되었습니다. 또한, 각 카세트에는 특이적인 barcode가 포함되어 있어 pooled library 상에서 다량의 유전자 기능 분석이 가능합니다.
- Heterozygous diploid deletion mutants 라이브러리의 경우 총 4,845개의 변이체로 구성되어 있고 이는 전체 게놈 (총 4,914개 ORF 포함)의 약 98%를 차지합니다.
- Homozygous haploid deletion mutants 라이브러리의 경우, 총 3,497개의 변이체로 구성되어 있습니다
각 균주들은 특이적인 barcode를 selection 마커 (KanMX4) 양쪽에 지니고 있으며, 이는 전체 균주 pool상에서 다량의 유전자 기능 분석 및 약물 타겟 스크리닝이 용이한 수단을 제공하고 있습니다. 또한, 각 개별 균주의 표현 형질을 분석함으로서 유전자의 기능을 규명하는데도 효과적으로 사용 가능합니다.
응용 분야
응용 분야 1. 항균제 대상 타겟 스크리닝
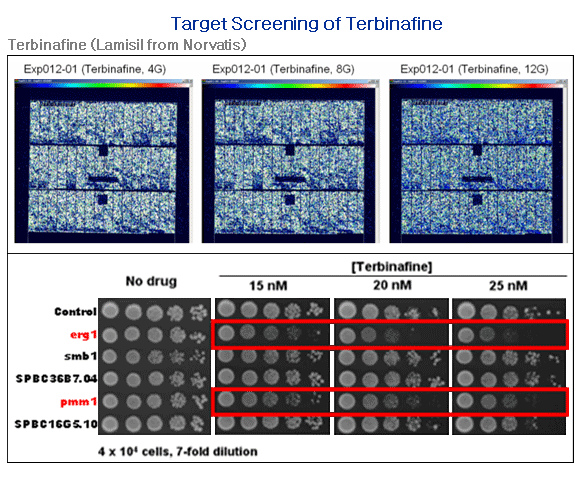
Figure 1. 항균제(Terbinafine)을 S.pombe deletion mutant library에 처리하여 기존에 알려진 타겟인 erg1 유전자를 확인하였고, 신규 타겟으로서 pmm1 유전자를 새로 발굴함.
 응용 분야 2. 고지혈증 치료제 대상 타겟 스크리닝
응용 분야 2. 고지혈증 치료제 대상 타겟 스크리닝
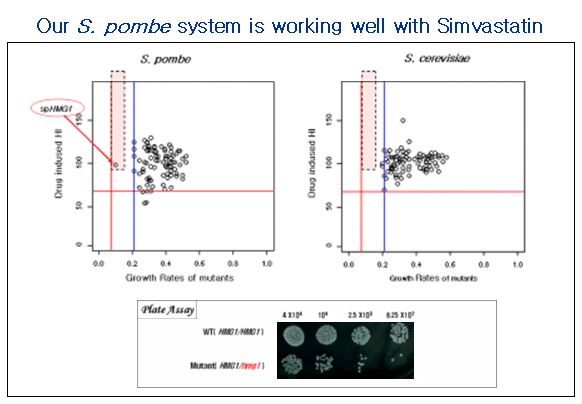
Figure 2. 고지혈증 치료제(simvastatin)의 타겟인 HMG1의 S.pombe ortholog인 hmg1 유전자를 타겟 스크리닝을 통해 확인함.