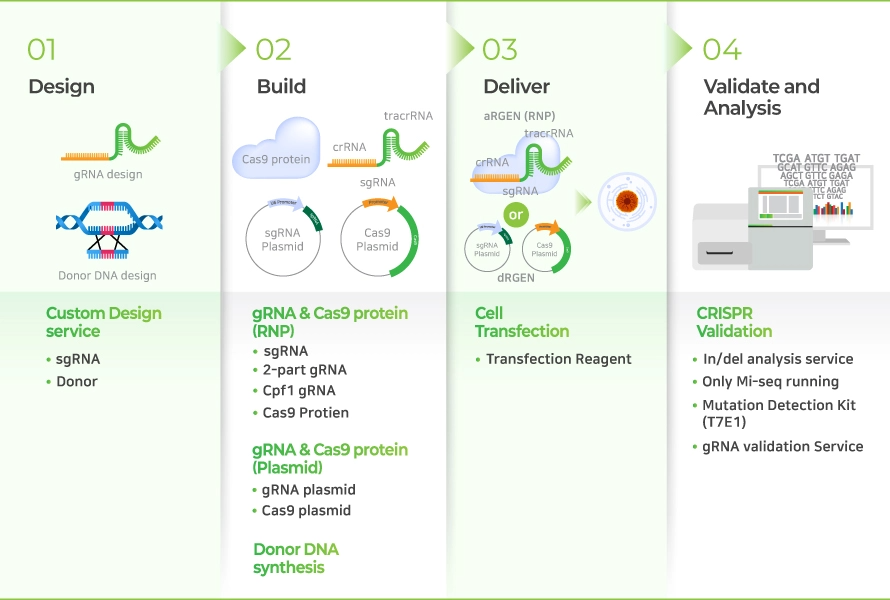
CRISPR (Clustered regularly interspaced short palindromic repeats)-Cas9 기술은 미생물의 적응면역 현상에서 기인한 유전자 편집 기술로, target DNA를 정밀하게 잘라낸 다음 자연적으로 DNA가 복구되도록 함으로써 보다 간단하고 효율적으로 유전자 조작을 가능하게 합니다. 이 시스템은 gRNA (guide RNA)와 Cas9 nuclease로 구성되며, RNA 유전자 가위 (RNA-guided engineered nucleases, RGENs) 기술이라고도 불립니다. gRNA는 target 서열에 상보적으로 결합하는 crRNA와 Cas9-binding을 위한 tracrRNA로 구성됩니다(Figure 1). gRNA는 Cas9을 target 부분으로 안내하는 역할을 하며, target DNA 서열과 상보적으로 결합합니다. Target DNA 서열 3’ 말단에는 PAM (Protospacer adjacent motif) 서열이 있어야 합니다. gRNA가 Cas9 nuclease와 복합체를 형성하고 target 서열에 특이적으로 결합하면, Cas9 nuclease는 PAM 서열을 인식하고 PAM의 3 bp upstream 부분에 이중 가닥 손상(Double Strand Break, DSB)을 일으킵니다(Figure 2).
CRISPR
CRISPR
Accurate Tool of CRISPR-Cas9 System
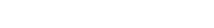
AccuTool™ | Accurate Tool of CRISPR-Cas9 System
CRISPR-Cas9 기술은 1세대 ZFN (Zinc Finger Nuclease), 2세대 TALEN (Transcription Activator-Like Effector Nuclease)보다 경제적이고 효율적인 강력한 3세대 Genome Editing Tool입니다. CRISPR-Cas9 시스템은 target DNA를 정확하게 인식하고 결합하는 guide RNA와 target DNA를 절단하는 Cas9 nuclease로 구성되어 DNA 이중 가닥을 손상시키고 이것을 복구하는 과정을 통해 유전자를 조작할 수 있습니다. 바이오니아는 유전자 편집 기술을 선도하고 있는 툴젠과 협력하여 런칭한 AccuTool™을 통해 gRNA 디자인부터 합성, Cas9 nuclease, Validation까지 Genome Editing 전반의 Total Solution을 제공합니다.
Workflows of CRISPR/Cas9
Workflows of CRISPR/Cas9
유전자 편집은 target 유전자의 gRNA를 디자인하면서 시작됩니다. gRNA는 Cas9 nuclease를 Target DNA 서열로 유도합니다.
NHEJ pathway를 이용하기 위해 실험에 맞는 gRNA와 Cas9 nuclease의 형태를 결정하고 합성합니다.
HDR pathway를 이용하기 위해 Donor DNA를 디자인 후 Donor synthesis를 합니다. 이후 gRNA와 Cas9 nuclease의 형태를 결정하고 합성합니다.
gRNA, Cas9 nuclease와 함께 target 세포에 transfection하여 정밀한 수정을 진행합니다.
유전자 편집 효율은 Mutation Detection Kit 또는 In/del analysis 서비스를 이용하여 Validation합니다.
CRISPR Cas9기술은 살아있는 유기체의 DNA를 정밀하게 수정할 수 있게 해주는 혁신적인 유전자 편집 기술입니다. Target DNA를 정확하게 인식하고 결합하는 guide RNA와 target DNA를 절단하는 Cas9 nuclease로 구성되어 DNA 이중 가닥을 손상시키고 이것을 복구하는 과정을 통해 유전자를 조작할 수 있습니다.
gRNA는 target 서열에 상보적으로 결합하는 crRNA와 Cas9-binding을 위한 tracrRNA로 구성됩니다. gRNA는 Cas9을 target 부분으로 안내하는 역할을 하며, target DNA 서열과 상보적으로 결합합니다. Target DNA 서열 3’ 말단에는 PAM (Protospacer adjacent motif) 서열이 있어야 합니다.
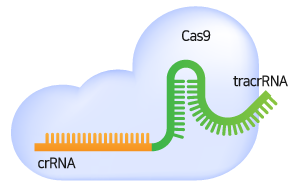
gRNA가 Cas9 nuclease와 복합체를 형성하고 target 서열에 특이적으로 결합하면, Cas9 nuclease는 PAM 서열을 인식하고 PAM의 3 bp upstream 부분에 이중 가닥 손상(Double Strand Break, DSB)을 일으킵니다.
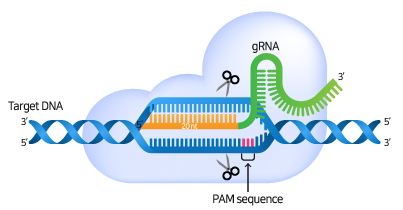
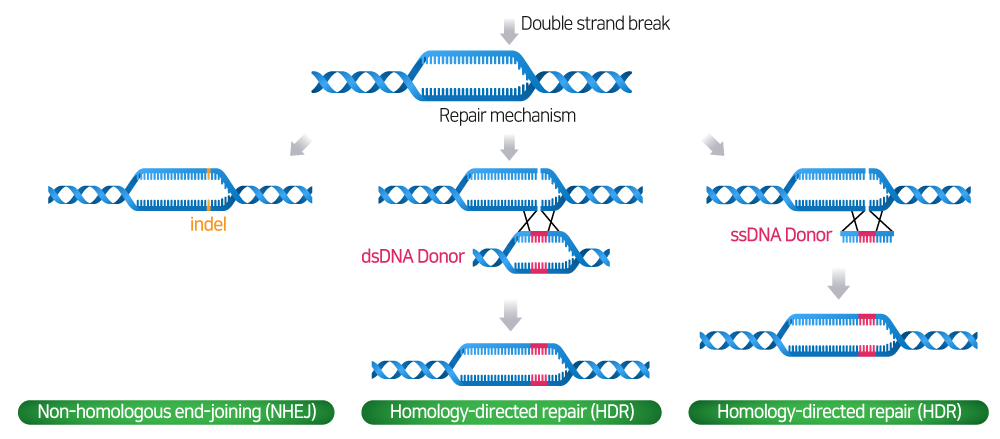
DSB로 인해 손상된 DNA를 복구하기 위하여 세포는 일반적으로 두 가지의 경로를 거치게 됩니다: Non-homologous end-joining (NHEJ) 또는 Homology-directed repair (HDR) pathway.
- NHEJ pathway: Donor template이 없는 경우 NHEJ pathway를 통해 DSB가 재조립되며, 유전자 coding region 내에 Indel (Insertion and deletion) 생성으로 인한 frameshifts 및 premature stop codon으로 유전자 Knock-Out이 만들어집니다.
- HDR pathway: HDR pathway는 Donor template에 의해 정밀하고 정확한 수정 및 새로운 유전자를 삽입할 수 있습니다.
CRISPR-Cas9 기술은 1세대 ZFN (Zinc Finger Nuclease), 2세대 TALEN (Transcription Activator-Like Effector Nuclease)보다 경제적이고 효율적인 강력한 3세대 Genome Editing Tool입니다.
| 기술 | 장점 | 단점 |
|---|---|---|
| ZFN | 높은 특이성 광범위한 적용 가능성 |
복잡한 설계 및 높은 비용 세포 독성 가능성 상대적으로 낮은 효율성 |
| TALEN | 높은 특이성 모듈화된 설계로 다양한 서열 타겟팅 가능 |
큰 크기로 인해 세포 내 전달 어려움 높은 비용 상대적으로 낮은 효율성 |
| CRISPR-Cas9 | 간편한 설계 높은 효율성 및 다중 유전자 타겟팅 가능 저렴한 비용 상대적으로 소요시간 빠름 다양한 생물 종과 조직에 적용 가능 |
ZFN과 TALEN에 비하여 면역반응이 있을 수 있음 |
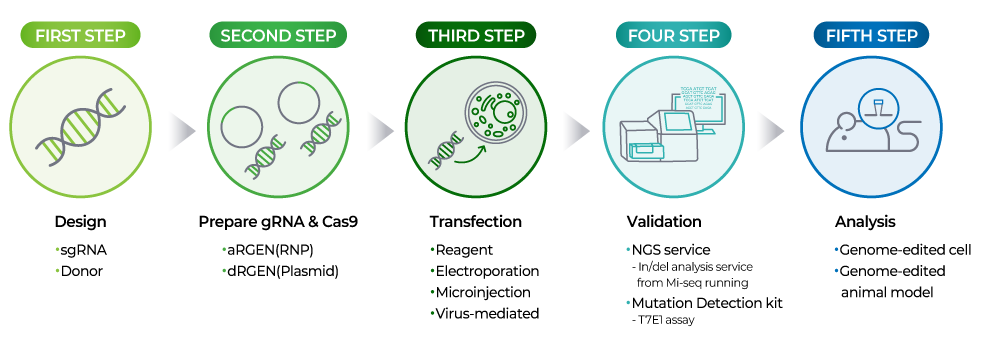
The CRISPR workflow involves the following steps:
- 1. gRNA 디자인: Knock out 및 knock in을 진행하려면 gRNA와 Donor 디자인이 필요합니다.
- 2. gRNA 합성: 디자인된 target sequence를 Plasmid 또는 ssRNA oligo 형태로 합성합니다.
- 3. cell에 도입 및 editing: 제작된 Cas9과 gRNA를 Target 세포에 transfection합니다.
- 4. Validation: Cell에서 gDNA prep 후 Mutation detection kit 또는 In/del analysis service를 통해 editing 효율을 확인합니다.
- 5. Analysis: 후속 실험을 진행합니다.
GC content는 40-80%, 길이는 18-20bp, sgRNA 서열 내 PAM 서열이 포함되지 않게 디자인하시는 것을 추천 드립니다. 사용하시고자 하는 Cas9의 PAM 서열을 원하시는 유전자에서 찾아 PAM site의 5’ upstream의 20nt로 디자인합니다.
sgRNA design이 어려우신 분들은 AccuTool™ Custom Design Service 이용하시면 됩니다.
해당 서비스는 높은 on-target efficiency, 낮은 off-target effects의 서열 3-4개를 디자인하여 제공합니다. 높은 편집 효율을 위해 sgRNA는 최소 2개 이상 디자인하여 실험에 사용하는 것을 추천 드립니다.
Off-target effects는 Cas9이 표적 부위가 아닌 다른 위치에서 작용하여 의도하지 않은 DNA를 절단하는 것을 의미합니다.
Off-target을 줄이기 위해서는 sgRNA 디자인이 가장 중요합니다. Off-target effects는 in silico tool을 이용하여 예측할 수 있으며 sgRNA 서열을 기반으로 계산됩니다.
sgRNA(dRGEN)과 Cas9(pRGEN)을 1:1 비율로 transfection하여 사용하시면 됩니다.
형광이 tagging된 plasmid를 사용하신다면 일차적으로 transfection 효율과 screening이 가능합니다.
CMV promoter와 EF1a promoter는 Cas9 단백질을 발현하는 promoter로 사용됩니다.
CMV Promoter:
- CMV promoter는 다양한 포유류 세포에서 높은 수준의 Cas9 단백질 발현을 유도하는 강력한 promoter입니다.
- 따라서 높은 Cas9 발현 수준이 필요한 경우 CMV promoter의 사용을 권장합니다.
- 하지만 CMV promoter의 강한 발현으로 인해 의도하지 않은 위치에서 표적을 벗어난 편집이 일어나거나 세포 독성을 유발할 수 있습니다.
EF1a Promoter:
- EF1a promoter는 여러 세포 유형에서 우수한 수준의 Cas9 발현을 제공하는 강력한 promoter입니다.
- CMV promoter와 달리 메틸화에 의한 유전자 silencing에 내성이 있기 때문에 안정적인 발현을 위해 적합한 promoter입니다.
- CMV promoter에 비해 표적을 벗어난 편집이 일어나거나 세포 독성을 유발할 가능성이 적습니다.
- 따라서 표적 외 효과 및 세포독성 최소화를 우선시하는 경우 EF1a promoter의 사용을 권장합니다.
promoter의 효과는 사용하는 특정 세포주에 따라 달라질 수 있습니다. 특정 세포주에 대한 권장 사항이 있는지 확인하려면 Cas9 플라스미드에 대한 문헌이나 기술 정보를 참조하여 선택하십시오.
sgRNA (single guide RNA)는 단일 RNA 분자로 custom-designed crRNA 서열과 scaffold tracrRNA 서열이 결합된 복합체를 말합니다.
2-part gRNA는 각각의 crRNA와 tracrRNA를 말합니다.
2-part gRNA를 실험에 적용하기 위해서는 crRNA와 tracrRNA를 hybridize하여 complex를 형성해야 합니다.
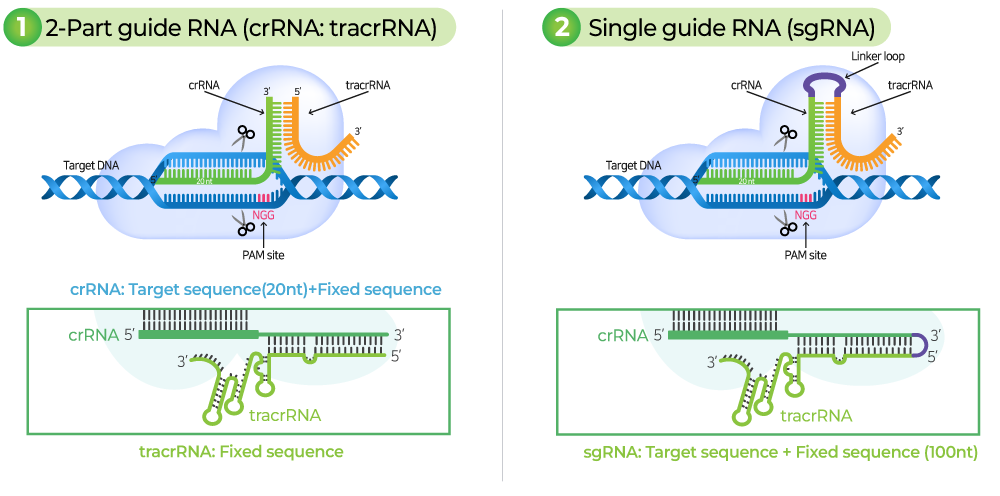
두 가지 모두 RNA oligonucleotide를 통해 합성이 되며 RNP 형태로 실험하기에 적합합니다. sgRNA (aRGEN)은 길이가 길기 때문에 2-part gRNA가 더욱 경제적이지만 sgRNA가 더욱 안정적입니다.
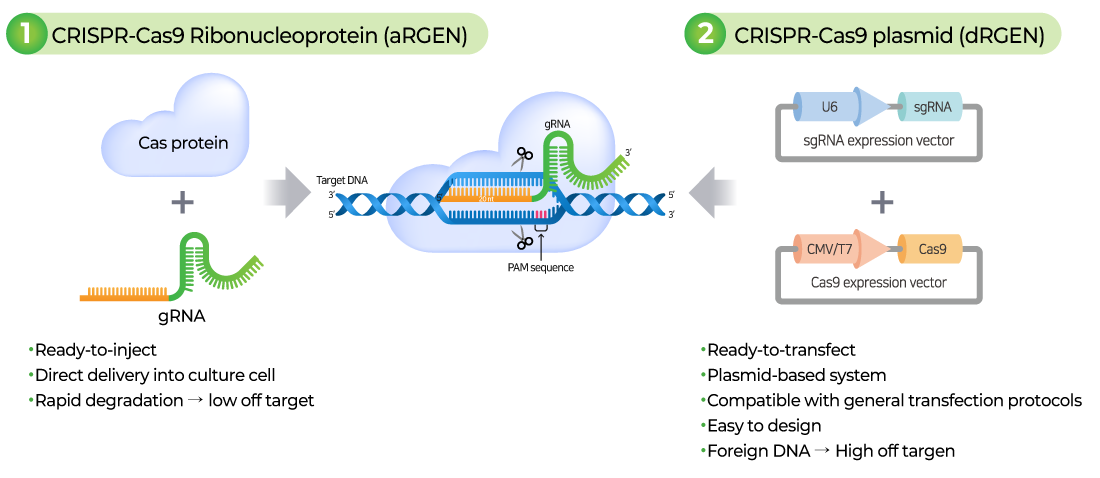
CRISPR-Cas9 유전자 편집을 위해 sgRNA와 cas9이 세포에 전달되는 형태로는 크게 두가지 RNP(ribonucleoprotein complex) 형태, Plasmid DNA 가 있습니다.
RNP (ribonucleoprotein complex):
- 구성: Cas9 protein + sgRNA (aRGEN)
- 전달: RNP complex -> transcription or translation 없이 직접 핵으로 전달 -> genome editing
- 장점:
- 빠른 gene editing 가능: 전사 과정이 필요하지 않아 Plasmid DNA에 비해 빠르게 실험이 가능합니다.
- 낮은 cytotoxicity: RNP는 protein과 sgRNA로 구성되어 있어 외부 DNA 전달이 필요하지 않으며 RNP는 빠르게 분해되어 낮은 cytotoxicity를 보입니다.
- 낮은 Off-target effect: 세포 내 잔여 시간이 짧고 proteases와 RNases에 의해 분해되어 낮은 off-target을 보입니다.
- in vitro, in vivo 모두 사용 가능
Plasmid DNA:
- 구성: Cas9 plasmid + plasmid sgRNA (dRGEN)
- 전달: 핵으로 이동 -> sgRNA, Cas9 transcription -> Cas9 translation -> RNP 형성 -> 핵으로 이동 -> genome editing
- 장점:
- 저렴한 비용: RNP에 비해 생산 및 조작 비용이 저렴합니다.
- 긴 보관 수명: Plasmid DNA는 RNP보다 더 안정적이고 긴 보관 수명을 갖습니다.
- 간편한 전달: 일반적인 형질전환 방법을 사용하기 때문에 간편하게 실험이 가능합니다.
- Screening 용이: selection marker 발현 가능하여 형질전환 후 세포내에 전달이 되었는지 확인할 수 있습니다.
- 단점:
- 느리고 효율성이 낮음: Cas9 발현 및 sgRNA 처리에는 시간이 걸리므로 RNP에 비해 편집 프로세스가 느리고 잠재적으로 효율성이 떨어집니다.
- 높은 Off-target effect: RNP에 비해 Cas9과 sgRNA가 세포내에 긴 기간 남아 있어 off-target 가능성이 높습니다.
- 높은 cytotoxicity: 외부 DNA가 전달되어 높은 cytotoxicity를 보입니다.
sgRNA의 평균 molecular weight는 32,212g/mol입니다.
ng으로 사용하시고자 하신다면 sheet에 기재된 분자량으로 하기 식으로 계산하시면 됩니다.
분자량 M.W. (g) X mole 수 (nmole) = sgRNA의 양 (ng)
| ssDNA Donor | dsDNA Donor |
|---|---|
|
|
Homology arm의 길이는 insert의 길이에 따라 달라집니다:
- 짧은 size insert (예: Point mutation): homology arm의 길이는 40nt-50nt를 추천 드립니다.
- 큰 size insert: homology arm의 길이는 300-800nt를 추천 드립니다.
Mutation detection Kit (T7E1):
T7 Endonuclease I (T7E1) 효소를 사용하여 빠르고 편리하게 KO 효율을 확인 가능한 Kit입니다. T7E1 효소는 mismatch를 식별하여 절단합니다. 해당 kit를 통해 단순하게 정량이 가능하지만 단일 뉴클레오티드의 정확한 분석은 불가능합니다.
In/del analysis service:
AccuCRISPR™ In/del analysis service 서비스는 NGS를 통해 유전체의 특정 부분을 분석하는 Targeted resequencing 방법을 이용합니다. 표적 sequencing 방법으로 CRISPR-Cas9을 통한 유전자 편집 효율을 빠르고 정확하게 정량화 가능합니다.
Crispr 효율은 다음과 같은 요인에 따라 결정됩니다:
sgRNA: 서열에 따라 효율이 달라지기에 sgRNA 서열 디자인이 중요합니다. 또한 2-3개의 sgRNA를 사용하시는 것을 추천 드립니다.
Delivery Method: Host cell 또는 organism에 맞는 전달 방법을 사용합니다.
- 전달 방법: Transfection reagent, viral-transduction, electroporation, Microinjection
Crispr component format: sgRNA와 Cas9이 cell로 전달되는 형태는 DNA, RNA 그리고 RNP(pre-formed ribonucleoprotein complex)가 있습니다. 각각의 형태에 따라 적절한 전달 방법을 선택하여야 합니다.
- DNA: 핵으로 이동 -> sgRNA, cas9 transcription -> cas9 translation -> RNP 형성 -> 핵으로 이동 -> genome editing
- RNA: Cas9 mRNA가 cytoplasm 안에서 translated -> Cas9 protein 형성 후 gRNA와 RNP complex 형성 -> 핵으로 이동 -> genome editing
- RNP: RNP complex -> transcription 또는 translation 없이 직접 핵으로 전달 -> genome editing
Cas9 variant 사용: 바이오니아에서는 SpCas9의 variant인 Sniper Cas9을 판매하고 있습니다. Sniper Cas9은 Spcas9 WT보다 낮은 off-target effect와 높은 specificity의 특징을 가지고 있습니다. Lee, J.K., Jeong, E., Lee, J. et al. Directed evolution of CRISPR-Cas9 to increase its specificity. Nat Commun 9, 3048 (2018).
I. Overview of Bioneer's AccuTool™ CRISPR-Cas9 Portfolio
A. Introduction to Comprehensive Solutions
Bioneer, operating in collaboration with ToolGen, presents its AccuTool™ portfolio as a comprehensive suite of products and services designed to facilitate CRISPR-Cas9 based gene editing. This portfolio aims to provide researchers with an integrated platform covering the essential stages of the gene editing workflow. The offerings span from the initial design and synthesis of guide RNA (gRNA) components to the provision of Cas9 nucleases in various formats, synthesis of donor DNA templates for knock-in applications, support for cellular delivery, and services for validating the outcomes of gene editing experiments. This integrated structure potentially offers researchers a streamlined pathway for acquiring necessary reagents and support from a single source, potentially simplifying experimental planning and execution compared to sourcing components individually.
B. Supported Gene Editing Modifications
The AccuTool™ services primarily support the two most common applications of the CRISPR-Cas9 system: gene knock-out and gene knock-in.
- Knock-out (Gene Disruption): This process aims to inactivate a target gene. It relies on the cell's natural Non-Homologous End Joining (NHEJ) DNA repair pathway, which is activated following the double-strand break (DSB) induced by the Cas9 nuclease at the gRNA-specified genomic locus. NHEJ is an error-prone repair mechanism that frequently introduces small insertions or deletions (indels) at the DSB site. These indels can cause frameshift mutations within the coding sequence, often leading to premature stop codons and the production of non-functional truncated proteins, effectively knocking out gene function. This approach does not require an exogenous DNA donor template.
- Knock-in (Gene Insertion/Replacement): This strategy enables the precise insertion or modification of DNA sequences at a targeted genomic location. It utilizes the cell's Homology-Directed Repair (HDR) pathway, which is generally less active than NHEJ but allows for high-fidelity repair using a template. Successful knock-in requires the co-delivery of the CRISPR-Cas9 components and a donor DNA template. This template contains the desired sequence (e.g., a point mutation, tag, reporter gene, or corrective sequence) flanked by sequences homologous to the genomic regions immediately surrounding the DSB site (homology arms). The HDR machinery uses this template to accurately repair the break, integrating the desired sequence into the genome.
The focus on these foundational gene editing techniques suggests that the AccuTool™ portfolio is primarily targeted towards researchers employing standard gene editing methodologies for functional studies, model generation, or screening purposes.
II. Core Components: Design and Synthesis
A. Guide RNA (gRNA)
Effective and specific targeting by the Cas9 nuclease is critically dependent on the design and quality of the gRNA. Bioneer offers services and products related to both gRNA design and synthesis.
1. Custom gRNA Design Service
Recognizing the importance of gRNA performance, Bioneer provides a custom design service. This service aims to identify optimal gRNA sequences targeting a user-specified locus. The design process considers key parameters influencing editing outcomes:
- On-target efficiency: The service proposes 3 to 4 candidate target sequences predicted to have the highest likelihood of successful cleavage at the intended site.
- Off-target effects: To minimize unintended cleavage at other genomic locations with sequence similarity, the design algorithm checks for potential off-target sites with up to 3 base pair mismatches compared to the intended target sequence. This explicit consideration of potential mismatches reflects an effort to enhance the specificity and reliability of the designed gRNAs, addressing a common concern in CRISPR experiments.
2. gRNA Synthesis Options
Bioneer provides gRNAs in multiple formats to accommodate different experimental requirements and delivery strategies.
- Chemically Synthesized sgRNA (aRGEN): Single guide RNAs (sgRNAs) are available as chemically synthesized RNA molecules. These are supplied in a lyophilized format, ready for reconstitution and complexing with Cas9 protein for Ribonucleoprotein (RNP) delivery. Bioneer provides positive control sgRNAs targeting human EGFP, CCR5, and HPRT1 genes. These sgRNAs are selected based on computational predictions for high on-target activity, offering researchers useful reference tools for optimizing CRISPR experiments. Custom synthesis based on user-provided or Bioneer-designed sequences is available.
- gRNA Expression Plasmids (dRGEN): Alternatively, gRNAs can be expressed intracellularly from plasmid vectors. Bioneer offers custom synthesis of gRNA sequences cloned into expression plasmids. Options include a basic vector utilizing the U6 promoter (dRGEN-U6-sgRNA) or a vector that co-expresses Green Fluorescent Protein (GFP) under a CMV promoter (dRGEN-U6-sgRNA-GFP-CMV). The GFP marker allows for monitoring of transfection efficiency via fluorescence microscopy. These plasmids are designed to recognize target sequences of 19-20 bp adjacent to a Protospacer Adjacent Motif (PAM) and are supplied ready for transfection. Positive control plasmids targeting EGFP, HPRT1, and CCR5 are also available.
- Other Formats: The service overview also mentions the availability of 2-Part gRNA and Cpf1 gRNA. The 2-part gRNA system consists of a crRNA and a tracrRNA. Since the tracrRNA sequence is fixed, this format offers a cost-effective solution for users who need to order multiple crRNA sequences.
Cpf1 (also known as Cas12a) is a CRISPR-associated endonuclease that utilizes a distinct guide RNA (gRNA) structure compared to Cas9. Bioneer offers custom synthesis services for Cpf1 gRNAs, including both AsCpf1 and LbCpf1 variants. This provides researchers with a convenient and reliable solution for Cas12a-based genome editing applications.
The availability of both chemically synthesized sgRNA (aRGEN) and plasmid-based gRNA expression vectors (dRGEN) provides researchers with flexibility. RNP delivery (using aRGEN and Cas9 protein) is often preferred for its rapid action, transient nature (reducing potential long-term off-target effects), and efficacy in difficult-to-transfect cells. Plasmid delivery (using dRGEN and pRGEN) can be simpler for laboratories routinely performing plasmid transfections and allows for the use of selection markers.
Table 1: gRNA Options Summary
| Product/Service | Format | Key Features |
|---|---|---|
| Custom gRNA Design Service | Service | 3-4 high-efficiency targets; Checks up to 3 bp mismatches for off-targets |
| sgRNA (aRGEN) | Chemically Synthesized RNA (Lyophilized) | Custom synthesis & positive controls. For RNP delivery |
| sgRNA Plasmid (dRGEN-Basic) | Plasmid DNA (Ready-to-transfect) | U6 promoter; Recognizes 19-20 bp target + PAM |
| sgRNA Plasmid (dRGEN-GFP) | Plasmid DNA (Ready-to-transfect) | U6 for sgRNA + CMV for GFP co-expression; Transfection monitoring |
B. Cas Nucleases
The Cas nuclease is the effector enzyme responsible for DNA cleavage. Bioneer offers several variants of the commonly used Streptococcus pyogenes Cas9 (SpCas9) and one ortholog, CjCas9, in different delivery formats.
1. Enzyme Variants
Bioneer provides a range of Cas9 variants catering to different experimental needs and applications:
- Streptococcus pyogenes Cas9 Wild-Type (SpCas9 WT): The standard enzyme for inducing DSBs.
- Streptococcus pyogenes Cas9 Nickase (SpCas9 D10A): Contains a mutation (D10A) inactivating one of the two nuclease domains, resulting in the generation of single-strand breaks (nicks) instead of DSBs. Paired nickases, targeting opposite strands in close proximity using two different gRNAs, can be used to generate a DSB with potentially increased specificity compared to WT Cas9.
- Streptococcus pyogenes Dead Cas9 (dCas9 D10A/H840A): Contains mutations in both nuclease domains (D10A and H840A), rendering it catalytically inactive for DNA cleavage. It retains its ability to bind DNA in a gRNA-programmed manner. dCas9 is primarily used as a platform for targeting functional domains (e.g., transcriptional repressors/activators, epigenetic modifiers) to specific genomic loci for applications like CRISPR interference (CRISPRi), CRISPR activation (CRISPRa), or epigenetic editing.
- Sniper Cas9: An engineered high-fidelity variant of SpCas9 designed to exhibit significantly reduced off-target activity compared to the WT enzyme, while maintaining high on-target efficiency. The development and characterization of Sniper Cas9 are detailed in a cited publication (Lee et al., Nat Commun, 2018). The provision of Sniper Cas9 directly addresses the critical concern of off-target mutations, offering a potentially safer and more precise option for applications demanding high specificity, such as therapeutic development or the generation of accurate disease models.
- Campylobacter jejuni Cas9 (CjCas9): A smaller Cas9 ortholog (protein size approx. 1000 amino acids vs. ~1370 for SpCas9; gene size ~2.9 kb for CjCas9). Its smaller size makes it particularly suitable for delivery methods with packaging size limitations, such as Adeno-Associated Virus (AAV) vectors. Bioneer offers CjCas9 in a plasmid format (pRGEN_CjCas9_CMV).
The availability of WT, Nickase, Dead Cas9, the high-fidelity Sniper Cas9, and the compact CjCas9 equips researchers with a versatile toolbox. This extends beyond simple gene knockout, enabling strategies like paired nicking for enhanced specificity, targeted transcriptional regulation or epigenetic modification using dCas9, and potentially facilitating in vivo delivery using viral vectors like AAV via CjCas9.
2. Delivery Formats
Cas9 nucleases are offered in two primary formats:
- Recombinant Protein: Purified Cas9 proteins (WT, Sniper, Nickase, Dead variants) are produced recombinantly in E. coli and feature a His-tag. They are supplied as a solution at a concentration of 1 mg/ml, accompanied by 10X Reaction buffer and 1X Dilution buffer. Recommended storage is between -70°C and -20°C. This format is intended for the preparation of RNP complexes by combining the protein with synthesized sgRNA (aRGEN) prior to delivery into cells. Bioneer provides specific quality control data for these proteins, including purity assessed as >95% by SDS-PAGE, absence of detectable contaminating RNase or DNase activity, and absence of protease activity. This detailed QC information provides assurance regarding the reagent's quality and suitability for sensitive applications.
- Expression Plasmids (pRGEN): Plasmids encoding human codon-optimized versions of Cas9 variants (WT, Sniper, Nickase, CjCas9) are available for intracellular expression following transfection. These pRGEN vectors offer choices for promoters (CMV/T7 or Ef1α) to drive Cas9 expression. Furthermore, several vectors incorporate selectable markers fused with fluorescent proteins (Puro-RFP or Hygro-EGFP), allowing for enrichment of transfected cells or tracking; basic vectors without markers are also available. Plasmids are supplied in quantities of 5 μg or 50 μg.
Table 2: Cas9 Nuclease Options Summary
| Cas9 Variant | Format | Key Specifications/Features |
|---|---|---|
| SpCas9 WT | Protein | 1 mg/ml solution; Incl. Buffers; >95% Purity; No DNase/RNase/Protease activity; His-tag. |
| SpCas9 WT | Plasmid | Human codon-optimized; Promoters: CMV/T7 or Ef1α; Markers: None, Puro-RFP, or Hygro-EGFP. |
| Sniper Cas9 | Protein | High-fidelity variant; 1 mg/ml solution; Incl. Buffers; >95% Purity; No DNase/RNase/Protease activity; His-tag. |
| Sniper Cas9 | Plasmid | High-fidelity variant; Human codon-optimized; Promoters: CMV/T7 or Ef1α; Markers: None, Puro-RFP, or Hygro-EGFP. |
| SpCas9 Nickase D10A | Protein / Plasmid | Nickase variant; 1 mg/ml solution; Incl. Buffers; >95% Purity; No DNase/RNase/Protease activity; His-tag. |
| SpCas9 Nickase D10A | Protein / Plasmid | Nickase variant; Human codon-optimized; Promoters: CMV/T7 or Ef1α; Markers: None, Puro-RFP, or Hygro-EGFP. |
| Dead Cas9 D10A/H840A | Protein | Nuclease-dead variant; 1 mg/ml solution; Incl. Buffers; >95% Purity; No DNase/RNase/Protease activity; His-tag. |
| CjCas9 | Plasmid | Small Cas9 ortholog (~2.9 kb); Vector: pRGEN_CjCas9_CMV. |
C. Donor DNA for Knock-in
Successful knock-in via HDR requires a donor DNA template. Bioneer offers services for both the design and synthesis of these templates.
1. Custom Donor Design Service
For researchers who need assistance in creating an effective donor template, Bioneer provides a Donor Design Service. Details are accessible via the general design service page. This service presumably helps optimize homology arm length and positioning of the insert for efficient HDR.
2. Donor Synthesis Options
Bioneer synthesizes custom donor templates in two main formats:
- Single-stranded DNA (ssDNA Donor / ssODN): These are chemically synthesized single-stranded oligonucleotides. Available synthesis scales cover lengths up to 2,000 nucleotides (nt), with specific ranges offered (e.g., 131-150 nt, 150-400 nt, 401-2,000 nt). ssDNA donors are often preferred for introducing point mutations or small insertions (e.g., tags). They are reported to potentially enhance knock-in efficiency, exhibit minimal cytotoxicity, and minimize random integration compared to dsDNA donors in certain experimental systems.
- Double-stranded DNA (dsDNA Donor): These templates are provided as double-stranded DNA, typically in 2-5 µg quantities. Although the exact format (e.g., plasmid, linear fragment) is not specified, dsDNA donors are generally used for inserting larger DNA fragments, as they can accommodate longer homology arms which may be beneficial for larger inserts. This format is described as commonly used and potentially having lower synthesis costs compared to long ssDNA donors.
The provision of both ssDNA and dsDNA donor synthesis options allows researchers to select the template format most appropriate for their specific knock-in experiment, considering the size of the intended insert and potential advantages related to efficiency and off-target integration associated with each type.
III. Delivery Systems and Support Services
Efficient delivery of CRISPR-Cas9 components into target cells is crucial for successful gene editing. Bioneer supports common delivery strategies and offers related reagents and services.
A. Delivery Strategies
The AccuTool™ platform is compatible with the two principal methods for delivering CRISPR components into cells:
- Ribonucleoprotein (RNP) Delivery: Involves pre-assembling purified Cas9 protein (e.g., AccuTool™ Recombinant Cas9) with chemically synthesized sgRNA (e.g., AccuTool™ aRGEN) in vitro to form RNP complexes, which are then introduced into cells (e.g., via electroporation or lipid-based transfection).
- Plasmid DNA Delivery: Involves transfecting cells with one or more plasmids encoding the Cas9 nuclease (e.g., pRGEN) and the gRNA (e.g., dRGEN).
The choice between RNP and plasmid delivery often depends on the specific cell type, and laboratory capabilities.
B. Transfection Reagents
Bioneer offers transfection reagents designed to facilitate the delivery of nucleic acids into cells. A link to the specific product ((https://www.bioneer.co.kr/20-k-7920.html)) is provided on the main service page, suggesting reagents optimized for use with their CRISPR system components are available.
C. Transfected Cell Service
For researchers seeking assistance with cell line generation, Bioneer offers the AccuTool™ Transfected Cell service, which delivers knock-out cell pools.
- Scope and Limitations: This service is specifically available for human adherent cell lines provided by the customer. The customer's cells must undergo and pass Mycoplasma testing by Bioneer prior to initiating the service. Additionally, the cells must express endogenous CCR5 or HPRT1 genes, which are used for positive control validation experiments. These prerequisites define the applicable scope of the service.
- Methodology: The service utilizes plasmid-based delivery, transfecting the customer's cells with plasmids encoding Cas9 and the customer-specified target gRNA. Bioneer optimizes the transfection conditions for the specific cell line. The reliance on plasmid delivery for this service contrasts with the availability of RNP components, suggesting a standardized workflow perhaps chosen for scalability or cost-effectiveness within the service context.
- Validation and Guarantees:Bioneer validates the efficiency of their optimized transfection and editing process using positive control gRNAs targeting CCR5 or HPRT1. They guarantee an editing efficiency (indel formation) of 40% or higher for these controls under the optimized conditions. However, a critical point is that the service does not guarantee the editing efficiency for the customer's specific target gRNA. The success of editing the intended target gene remains dependent on the intrinsic activity of the gRNA chosen by the customer. This service does not perform gRNA screening; researchers needing to identify highly active gRNAs are directed to the separate gRNA Validation service. This distinction clarifies that the service guarantees the process execution but not the outcome for a novel target.
- Deliverables: Customers receive two vials of the generated knock-out cell pool (1×106 cells/ml per vial), the sgRNA plasmid DNA (2 µg) used for the transfection, and a report detailing the transfection efficiency test results and the measured editing efficiency data for both the positive control and the customer's target.
IV. Pre-packaged CRISPR Kits
Bioneer offers pre-configured kits bundling necessary components for specific CRISPR applications, potentially simplifying ordering and experimental setup.
A. AccuTool™ CRISPR-Cas9 Starter Kit
This kit is designed as an introductory package for researchers initiating CRISPR experiments, particularly those utilizing plasmid-based approaches.
- Components: The kit includes three custom sgRNA plasmids (dRGEN, allowing for testing multiple gRNAs for a target), two vials of Cas9 expression plasmid (pRGEN-Cas9-CMV/T7), a Mutation Detection Kit (T7E1) for initial validation of editing, a positive control dRGEN plasmid (targeting EGFP, CCR5, or HPRT1), corresponding primers for the positive control locus, and a user manual.
- Applications: It provides a convenient starting point for performing gene knock-out experiments using plasmid delivery. The inclusion of multiple custom sgRNAs facilitates screening for effective guides, and the T7E1 kit allows for rapid assessment of editing activity.
B. AccuTool™ Safe Harbor Knock-In Kits (AAVS1/Rosa26)
These kits are specialized for targeted gene insertion into well-characterized genomic "safe harbor" loci, aiming for stable and predictable transgene expression. Separate kits are available for targeting the human AAVS1 locus and the mouse Rosa26 locus.
- Components: Each kit contains a pre-designed sgRNA plasmid (dRGEN) targeting the specific safe harbor site (AAVS1 or Rosa26), a Cas9 expression plasmid (pRGEN-Cas9-CMV/T7), a donor plasmid vector containing homology arms for the respective safe harbor locus and likely a cloning site for inserting the user's gene-of-interest (GOI), primers for T7E1 analysis of the target locus, and a user manual.
- Purpose and Applications: Safe harbor loci like AAVS1 and Rosa26 are genomic locations known to permit stable, robust expression of integrated transgenes without causing apparent detrimental effects on the host cell, such as insertional mutagenesis or gene silencing often associated with random integration. These kits provide optimized reagents (gRNA, donor vector) to facilitate efficient targeted integration of a GOI into these reliable genomic sites. Applications include creating stable cell lines for gene function studies, therapeutic research requiring consistent gene expression, and lineage tracing experiments.
The offering of both a general Starter Kit and specialized Safe Harbor Kits demonstrates an effort to cater to different levels of experimental complexity and user needs. The Starter Kit lowers the barrier for entry into plasmid-based CRISPR, while the Safe Harbor Kits provide a sophisticated solution for researchers requiring reliable and stable transgene expression, addressing a common challenge in genetic engineering.
V. Validation and Quality Control
Ensuring the quality of reagents and validating the outcomes of gene editing are critical aspects of CRISPR workflows. Bioneer provides QC data for components and offers tools and services for downstream validation.
A. Component Quality Control
Specific quality control measures are reported for key components:
- Recombinant Cas9 Protein: Assessed for purity (>95% by SDS-PAGE) and tested for the absence of contaminating nuclease (RNase, DNase) and protease activities.
- Positive Control sgRNAs (aRGEN): Selected based on computational predictions for high on-target activity and have passed quality control (QC) using MALDI-TOF mass spectrometry.
- Positive Control dRGEN Plasmids: Supplied with corresponding primer sets for validation.
B. Post-Editing Validation Services & Tools
Bioneer offers several methods to assess the efficiency and outcomes of gene editing experiments:
- Mutation Detection Kit (T7E1): This kit utilizes the T7 Endonuclease I enzyme, which recognizes and cleaves heteroduplex DNA formed between wild-type and indel-containing DNA strands following PCR amplification of the target locus. Gel electrophoresis of the cleavage products provides a relatively rapid, qualitative, or semi-quantitative indication of whether editing has occurred (presence/absence and intensity of cleavage bands). This kit is included in the Starter Kit and can be purchased separately.
- In/del Analysis Service (NGS-based): This service provides a more quantitative and detailed assessment of editing outcomes using Next-Generation Sequencing (NGS). It involves targeted deep sequencing (targeted resequencing) of the genomic region surrounding the CRISPR target site. Bioneer guarantees specific data quality metrics for this service, including a Phred quality score (Q30) exceeding 80% and a total read depth of more than 10,000 reads. These metrics ensure high base-calling accuracy and sufficient sampling depth for reliable quantification of various indel types and their frequencies. The service delivers an analysis report designed for clarity, along with the raw sequencing data. The emphasis on specific quality thresholds (Q30 > 80%, >10k reads) underscores a commitment to providing robust, high-resolution data suitable for rigorous analysis of editing efficiency and patterns.
- gRNA Validation Service: This is described as an experimental service to determine the editing efficiency of specific gRNAs. It is positioned as the appropriate avenue for screening and identifying highly active gRNAs, distinct from the Transfected Cell Service which does not guarantee customer target efficiency. The sequence of guide RNA (gRNA) plays a critical role in the efficiency of CRISPR experiments. Therefore, careful design of gRNA prior to conducting CRISPR-based gene editing is essential. Although some gRNA sequences may be predicted to have high activity based on theoretical models, actual experimental outcomes can vary significantly. As a result, it is crucial to validate gRNA efficiency experimentally to select the most effective sequence.
This service utilizes a plasmid-based delivery method to transfect the customer's cells with 4 gRNA sequences provided by the customer. The gene editing efficiency of each gRNA is evaluated using either a mutation detection kit or In/Del analysis. Through this process, customers can identify the gRNA sequence that exhibits the highest editing efficiency among the four candidates provided.
The availability of different validation methods, from the rapid T7E1 assay to the quantitative NGS analysis, provides researchers with a tiered approach. They can select the validation strategy that best fits their experimental stage, throughput needs, required level of detail, and budget – ranging from initial screening to precise characterization of edited alleles.
Table 4: Validation Methods Summary
| Method/Product | Principle | Output/Metrics | Format |
|---|---|---|---|
| Mutation Detection Kit (T7E1) | T7E1 cleavage of heteroduplex DNA | Qualitative/Semi-quantitative indel detection (gel electrophoresis) | Kit |
| In/del Analysis Service (NGS) | Targeted deep sequencing | Quantitative indel frequency; Q30 >80%, >10k reads | Service |
| gRNA Validation Service | Experimental editing efficiency test | Efficiency data for 4 gRNAs (report) | Service |
VI. Conclusion
A. Summary of Capabilities
Based on the reviewed technical information, Bioneer's AccuTool™ portfolio represents an integrated platform for CRISPR-Cas9 gene editing. It provides a comprehensive range of products and services encompassing critical steps from initial design (gRNA, donor templates) through reagent provision (diverse Cas9 variants and gRNA formats, donor DNA) and delivery support (transfection reagents, transfected cell service) to downstream validation (T7E1 kit, NGS analysis, gRNA validation service). This integrated offering aims to support researchers throughout the gene editing workflow.
B. Highlighted Technical Features
Several technical features stand out within the AccuTool™ portfolio. The availability of the high-fidelity Sniper Cas9 addresses the crucial need for minimizing off-target effects. Flexibility is provided through multiple delivery formats for both gRNA (synthesized RNA, expression plasmids) and Cas9 (recombinant protein, expression plasmids), accommodating both RNP and plasmid-based workflows. For knock-in experiments, the support for both ssDNA and dsDNA donor template synthesis caters to different insert sizes and experimental strategies. The provision of specialized Safe Harbor Knock-in Kits (AAVS1, Rosa26) offers a targeted solution for achieving stable and reliable transgene expression. Finally, the validation options include both rapid screening (T7E1) and rigorous quantitative assessment via NGS, with defined quality metrics for the sequencing service.
C. Overall Impression
The AccuTool™ CRISPR-Cas9 solutions appear designed to serve a broad spectrum of researchers engaged in gene editing. The portfolio includes entry-level options like the Starter Kit, suitable for those new to the technology or preferring plasmid workflows, as well as advanced components and services like Sniper Cas9, Safe Harbor kits, and quantitative NGS validation, catering to more demanding applications requiring high precision, reliability, and detailed characterization. The emphasis on component quality control (e.g., for Cas9 protein) and the provision of multiple validation tools suggest a focus on enabling robust and reproducible gene editing experiments. While the portfolio is comprehensive, potential users should note that specific technical details for some services, were not available in the reviewed materials and would require direct communication with Bioneer for full evaluation.